Research software
Easy-DHPSF
Automated processing of double-helix (DH) microscope images of single molecules (SMs) streamlines the protocol required to obtain super-resolved three-dimensional (3D) reconstructions of ultrastructures in biological samples. Easy-DHPSF is a suite of MATLAB subroutines, bundled with an easy-to-use graphical user interface, that facilitates 3D localization of single emitters (e.g. SMs, fluorescent beads, or quantum dots) with precisions of tens of nanometers in multi-frame movies acquired using a wide-field DH epifluorescence microscope. The algorithmic approach is based upon template matching for SM recognition and least-squares fitting for 3D position measurement, both of which are computationally expedient and precise. Once calibrated, the algorithm can fit 15-30 molecules per second on a 3 GHz Intel Core 2 Duo workstation, thereby producing a 3D super-resolution reconstruction of 100,000 molecules over a 20×20×2 μm field of view (processing 128×128 pixels × 20000 frames) in 75 min.
Features
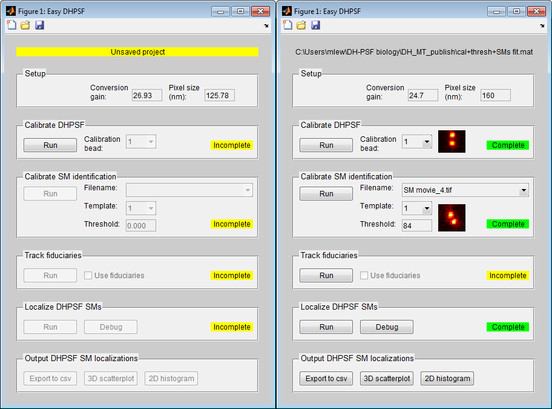
- Double-Gaussian estimator is calibrated via an axial scan of bright immobile fluorescent emitters (e.g. beads).
- Template-matching and double-helix recognition algorithm is calibrated via setting an image correlation threshold against single-molecule data.
- Tiff stacks of SM images are analyzed using template matching followed by double-Gaussian fitting to extract estimates of the molecule positions.
- The motion/drift of fiduciaries (e.g. fluorescent beads) during an experiment can tracked and removed form SM localization data.
- 3D localization data can be exported to .csv format for post-processing and visualization.
- Basic 3D scatterplot and 2D z-projection histogram visualizations are included.
Download
Easy-DHPSF project page at Google Code
Documentation
M. D. Lew, A. R. S. von Diezmann, and W. E. Moerner, "Easy-DHPSF open-source software for three-dimensional localization of single molecules with precision beyond the optical diffraction limit," Protocol Exchange (2013). [Journal, PDF]